FDA Issues Guidance on Submitting NGS Data for Antiviral Drugs
On July 18, the FDA issued final guidance on submission of next-generation sequencing (NGS) data from resistance assessments performed in the development of antiviral drug products. The new Technical Specifications document (Tech Doc) provides nonbinding recommendations on acceptable NGS platforms and the types of information the FDA is looking for sponsors to submit, including NGS protocols and data analysis methods. What’s At Stake Developers of antiviral drugs and related diagnostic tests use NGS to perform sequence-based resistance analysis. But while NGS allows for analysis of individual viruses within a viral population, the data it generates is highly complex and hard to validate, especially since there are currently no standardized bioinformatics analysis approaches for analyzing these large datasets. The significance of the new Tech Doc is in helping sponsors submit next generation nucleotide sequence analysis procedures and data in support of resistance assessments for the development of antiviral drugs and related tests. Specifically, the Tech Doc provides crucial guidance on five key issues. 1. Which NGS Platforms Are Acceptable The Tech Doc says that the FDA Division of Antiviral Products (Division) will accept nucleotide sequencing data generated from most standard NGS platforms as long as the sponsor submits: The appropriate details […]
On July 18, the FDA issued final guidance on submission of next-generation sequencing (NGS) data from resistance assessments performed in the development of antiviral drug products. The new Technical Specifications document (Tech Doc) provides nonbinding recommendations on acceptable NGS platforms and the types of information the FDA is looking for sponsors to submit, including NGS protocols and data analysis methods.
What's At Stake
Developers of antiviral drugs and related diagnostic tests use NGS to perform sequence-based resistance analysis. But while NGS allows for analysis of individual viruses within a viral population, the data it generates is highly complex and hard to validate, especially since there are currently no standardized bioinformatics analysis approaches for analyzing these large datasets. The significance of the new Tech Doc is in helping sponsors submit next generation nucleotide sequence analysis procedures and data in support of resistance assessments for the development of antiviral drugs and related tests. Specifically, the Tech Doc provides crucial guidance on five key issues.
1. Which NGS Platforms Are Acceptable
The Tech Doc says that the FDA Division of Antiviral Products (Division) will accept nucleotide sequencing data generated from most standard NGS platforms as long as the sponsor submits:
- The appropriate details for the sequencing platform;
- The protocols used for sample preparation;
- The raw NGS data in fastq format; and
- The methods used to analyze the data.
Sponsors should communicate with the Division early in the process and submit a mock NGS dataset before any formal submissions to verify that the appropriate data formats and processes are acceptable.
2. Information about NGS Protocol
Sponsors should also submit a detailed NGS protocol that includes six design elements:
- A description of the subjects, study time points and sample matrices to be analyzed;
- A description of the NGS platform and all associated performance characteristics;
- Target gene region name(s) and size(s) to be analyzed;
- A description of the general analysis strategy;
- The coverage level to be attempted (the Tech Doc recommends a target for coverage of greater than 5,000 reads while recognizing that this may not be possible for samples with lower viral loads); and
- A description of the approach used to identify, filter or process sequencing errors.
3. Frequency Tables
Sponsors should provide a frequency table reporting all amino acid substitutions that differ
from baseline at frequencies greater than or equal to 1%. The Tech Doc includes a model frequency table:
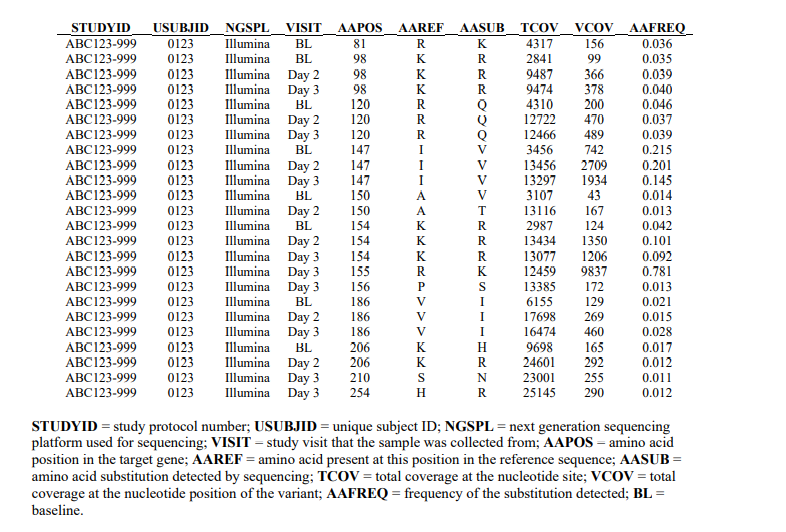
4. Information about Sample Preparation
Noting that the key to reliable sequencing results is sample preparation and ensuring that the sample sequenced is representative of the population analyzed, the Tech Doc recommends that sponsors list their methods for:
- Extracting nucleic acids from samples;
- Purifying viral sequences from contaminating background nucleic acids;
- Concentrating viral nucleic acids, including the estimated target copy number input for reverse transcription polymerase chain reaction (RT-PCR) (viral RNA) or PCR (viral DNA) reactions for each sample;
- Denaturing secondary structure;
- Generating double stranded DNA (dsDNA), including a description of the primers;
- Purifying dsDNA for sequencing;
- NGS library preparation; and
- Adding barcodes for multiplexing (if applicable).
Additional Standards for PCR Amplification
The Tech Doc says that any protocol that uses a PCR amplification step before NGS should provide evidence that amplifications are representative of the target population and that minor variants would still be present in the NGS data. The FDA recommends using approaches that correct for PCR resampling bias and (RT-)PCR and sequencing error, such as complementary DNA barcoding.
5. Information about Data Analysis & Reporting Results
Submissions of sequence data must include a thorough description of the analysis pipeline used
to analyze the sequencing dataset and the raw sequence information so that the Division can
conduct an independent analysis of the data. That would include the following information:
- Summary statistics for each sequence run, including total number of reads sequenced per sequence run, quality scores and average length of reads;
- A description of how sequence barcodes were processed;
- Contig and mapping reports—the Tech Doc recommends two data analysis approaches and establishes standards for each: i. mapping of short reads to a reference sequence; or ii. de novo assembly of short reads to assemble contigs.
6. Which Data File Types Are Acceptable
The Tech Doc calls on sponsor to provide all of the raw NGS data from each sequence run in the fastq format, which may also include an assembled read mapping in .fas, .ace, .sam, or .bam formats along with the appropriate reference sequences and accession numbers used for any reference mappings. For reference mapping to a baseline sample or gene of interest, the sponsor should provide the baseline or reference consensus sequence and state how this sequence was derived. For de novo assemblies, the sponsor should provide all contigs greater than 200 nucleotides in the fastq format.
The raw NGS data and frequency tables should and frequency tables should be sent to the Division on a secured, portable hard drive following the FDA Transmitting Electronic Submissions Using eCTD Specifications (April 2019) guidelines. Don't include any additional files on the hard drive, e.g., .exe extensions, or the submission may be rejected.
Takeaway: The new Tech Doc should make it easier to use NGS data to secure approval for antiviral drugs and companion tests. Although it's final guidance, the FDA is still accepting public comments on the document.
Subscribe to view Essential
Start a Free Trial for immediate access to this article