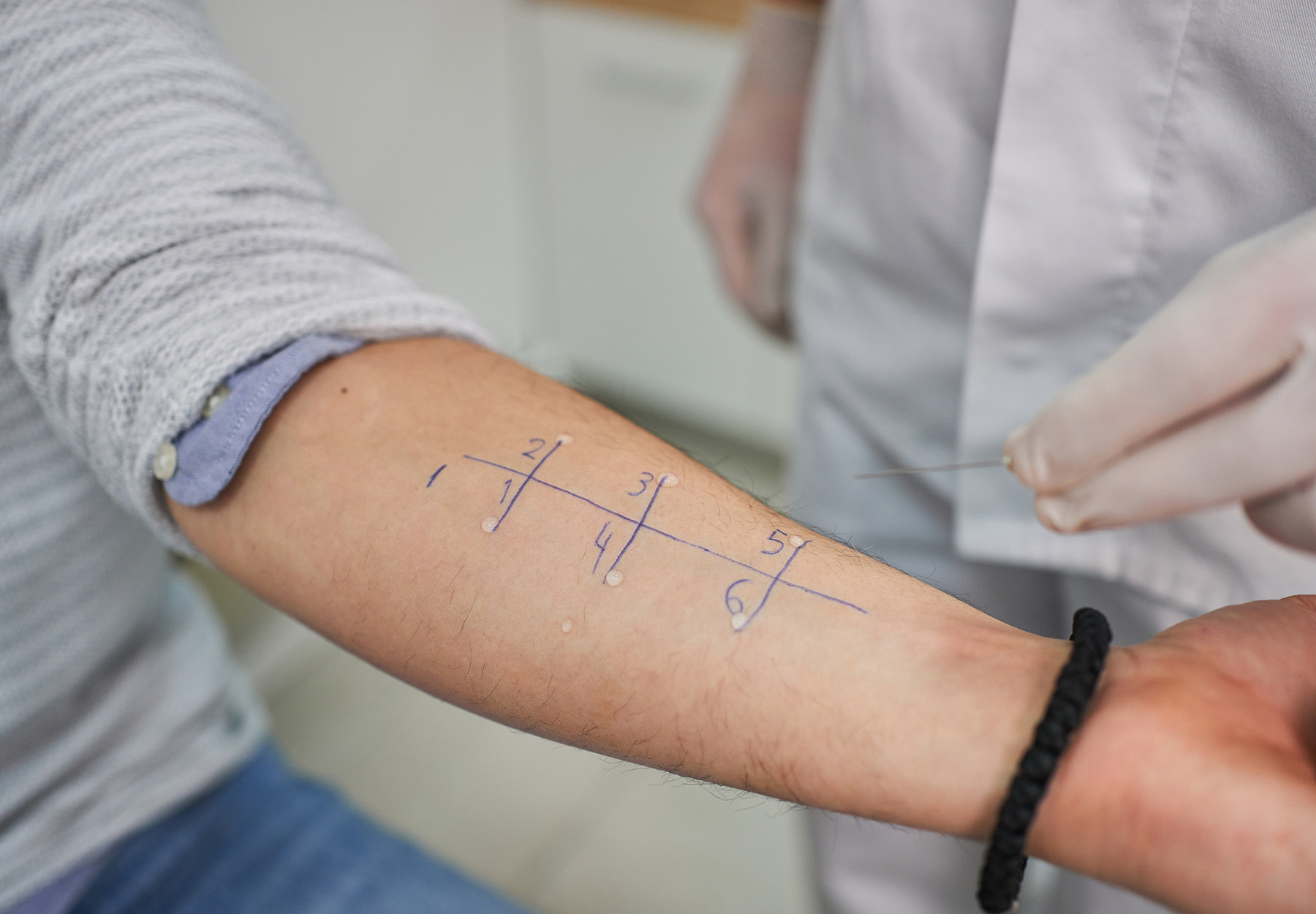
Diagnosing allergies is difficult and cumbersome. Current skin-prick tests are painful, time-consuming, and carry the chance of the patient having a dangerous allergic reaction. But a new assay from researchers at the University of Bern and Inselspital, Bern University Hospital, both in Switzerland, could make diagnosing allergies much easier, as well as give a better idea of how successful treatment for the allergy will be.
The assay, recently described in the Journal of Allergy and Clinical Immunology, focuses on the most common type of allergy, type I, which includes asthma, rhinitis, and eczema. These allergies occur when the immune system produces immunoglobulin E (IgE) antibodies in response to an allergen. These IgE antibodies bind to immune cells called mast cells, which release chemicals that cause an allergic reaction, such as a sore throat or rash. The researchers used these mast cells as the basis of their assay.
The researchers genetically engineered a cell line from the bone marrow of mice transgenic for the human high-affinity IgE receptor (FcεRIα), generating mature mast cells that behave similarly to how human cells would in an allergic reaction. They sensitized those cells with blood serum from allergic patients, then presented the cells with certain allergens, finding that their mast cell assay can:
- Identify culprit allergens
- Accurately measure total IgE levels
- Longitudinally monitor allergen-specific immunotherapy
- Possibly determine when that immunotherapy has started to work
In addition, the researchers showed that the test could potentially be used to analyze large volumes of samples, showing a proof of concept for a high-throughput screening application based on fluorescent cell barcoding with the engineered mast cells.
“Our results indicate that this novel mast cell assay could represent a valuable tool to support clinicians in the identification of IgE-mediated allergies and in the quantification of treatment efficacy as well as duration of therapeutic response,” the researchers conclude.